1. Upload
Drop in your 3D volume
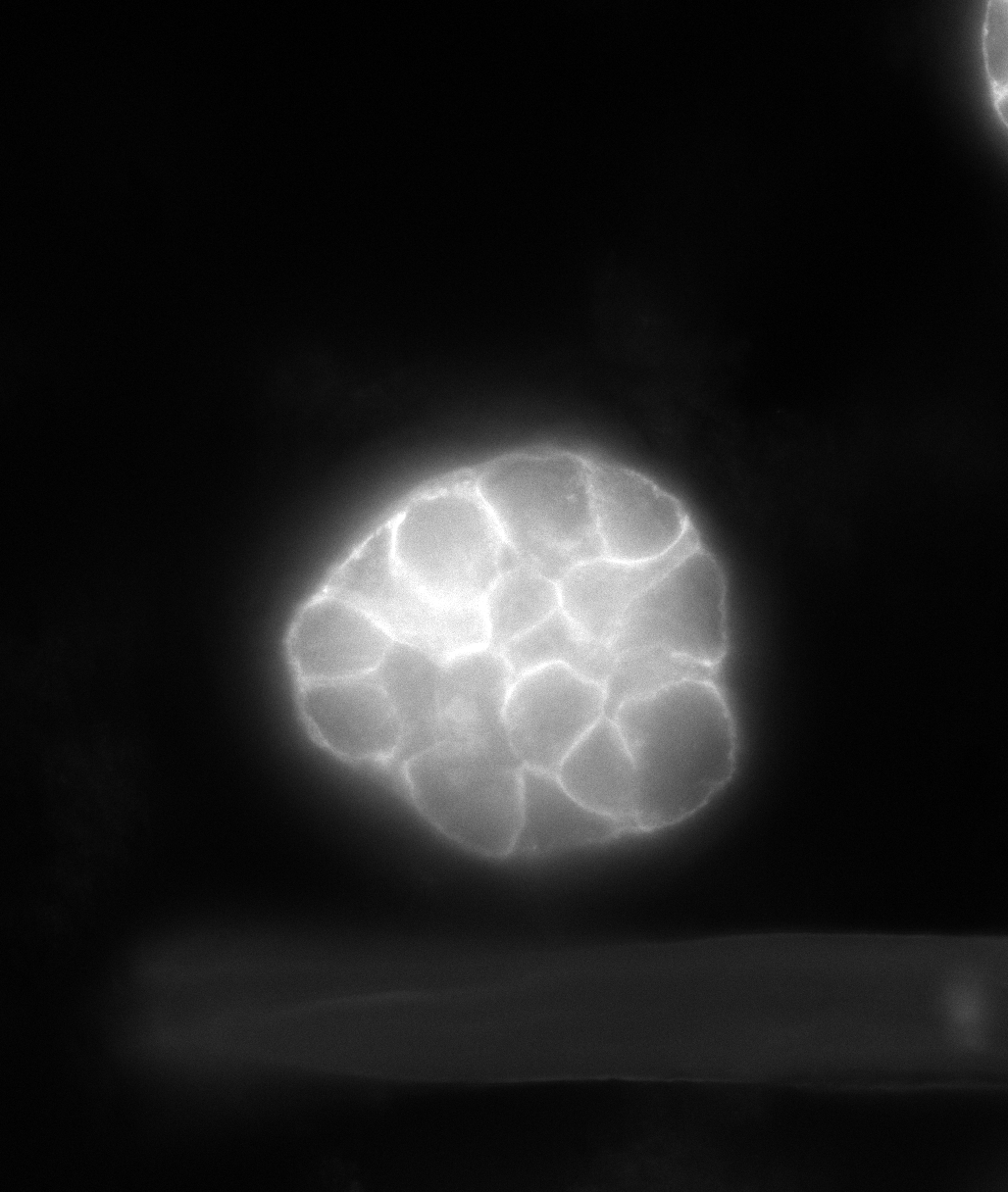
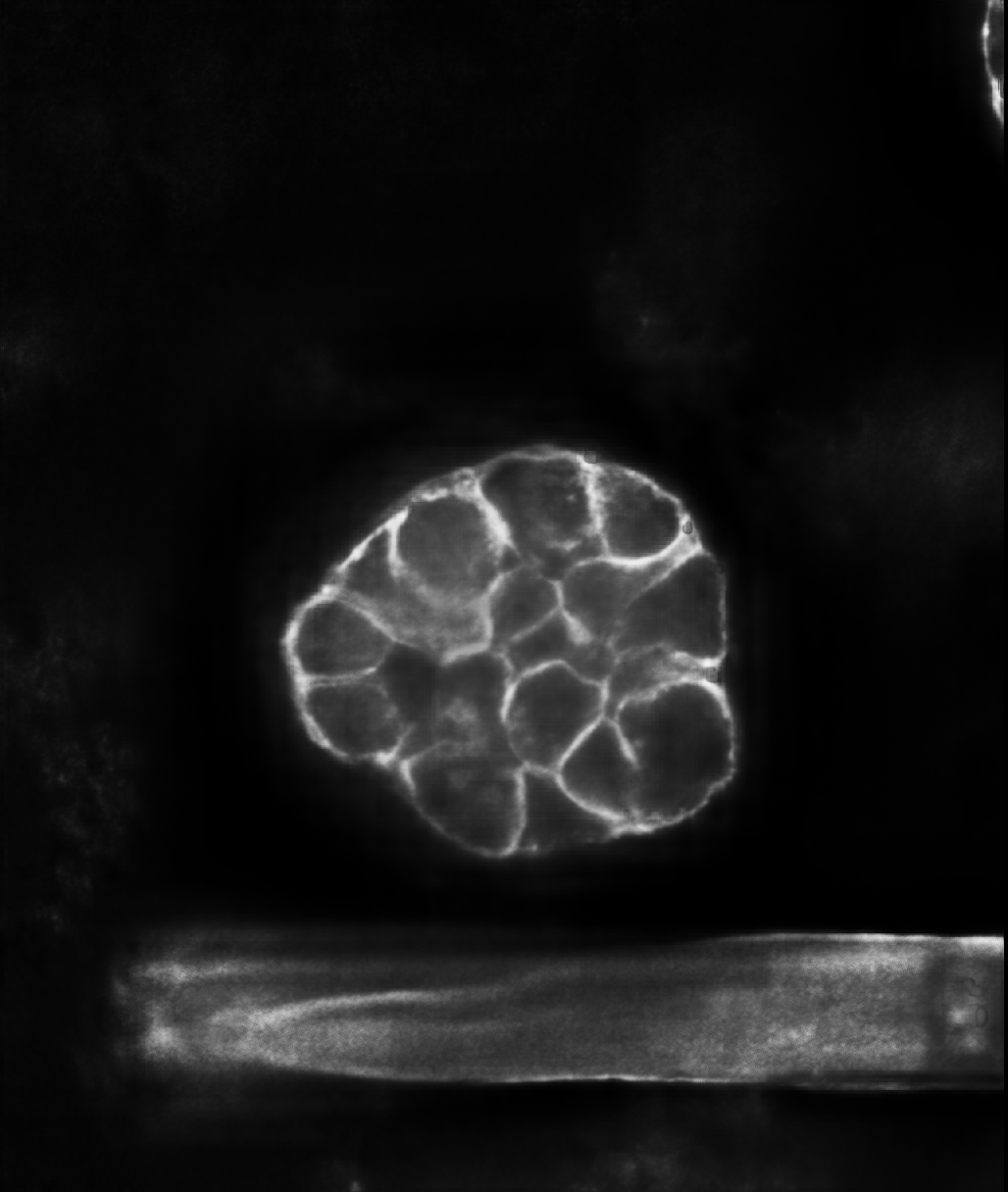
Upload 3D Volume directly into the browser and select model Linear(b-catenin, Filamentous) or Blob(DAPI).
No local installs, drivers, or Python environments to manage.
- Supports fluorescence Z-stacks
- Secure, session-based processing
- Works with data from low-cost microscopes
2. Enhance
AI-powered deconvolution
Our custom 3D AI model denoises, sharpens, and deconvolves while preserving delicate boundaries and
small structures critical for quantification.
- 3D context across the full volume
- Optimized for fast inference on standard GPUs
- Trained to avoid hallucinating structures
3. Export
Visualize and download
Scroll through enhanced slices, compare input vs output, and download the deconvolved output for
segmentation, tracking, or downstream AI tasks.
- Side-by-side review for quick QC
- Compatible with napari, Fiji, and Python workflows
- Ideal for organoid screens and assay readouts